Characterization and Development of EST-SSR Markers from Transcriptome Sequences of Chrysanthemum (Chrysanthemum ×morifolium Ramat.) in: HortScience Volume 54 Issue 5 (2019)
The genetic diversity and relationships of cauliflower (Brassica oleracea var. botrytis) inbred lines assessed by using SSR markers | PLOS ONE

Development of unigene-derived SSR markers in cowpea (Vigna unguiculata) and their transferability to other Vigna species

Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports
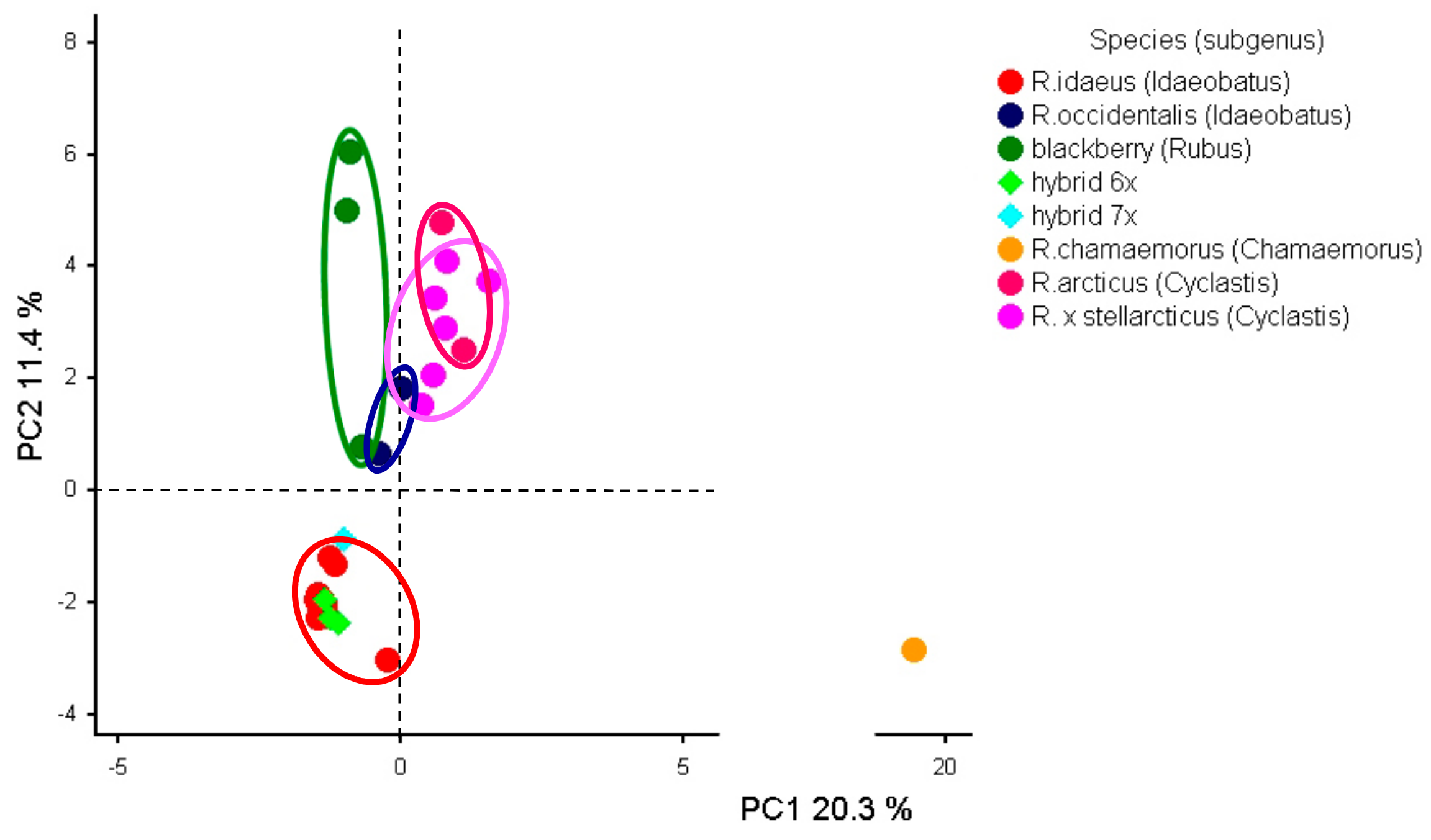
Genes | Free Full-Text | Transferability and Polymorphism of SSR Markers Located in Flavonoid Pathway Genes in Fragaria and Rubus Species
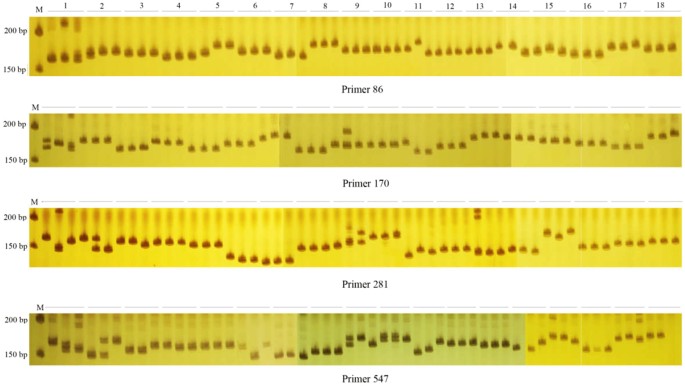

Cross-species transferability of EST-SSR markers developed from the transcriptome of Melilotus and their application to population genetics research | Scientific Reports

Comprehensive functional analysis and mapping of SSR markers in the chickpea genome (Cicer arietinum L.) - ScienceDirect
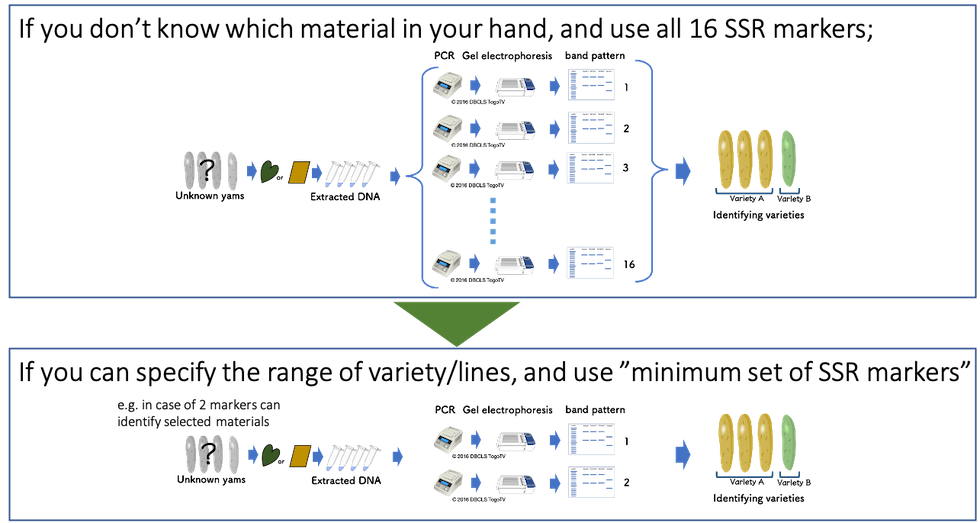
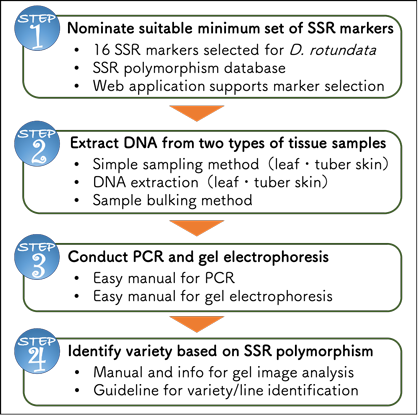
Yam Variety Identification Toolkit:Step 1 | Japan International Research Center for Agricultural Sciences | JIRCAS
![The newly developed genomic-SSR markers uncover the genetic characteristics and relationships of olive accessions [PeerJ] The newly developed genomic-SSR markers uncover the genetic characteristics and relationships of olive accessions [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/8573/1/fig-2-2x.jpg)
The newly developed genomic-SSR markers uncover the genetic characteristics and relationships of olive accessions [PeerJ]
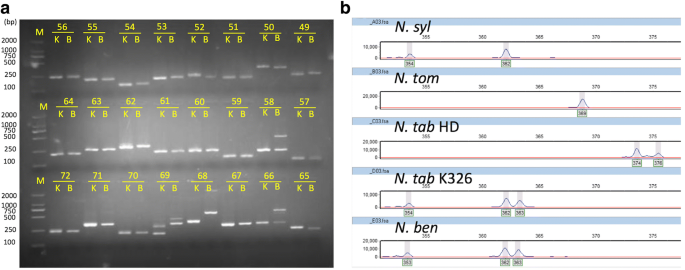
Comparative genome-wide characterization leading to simple sequence repeat marker development for Nicotiana | BMC Genomics | Full Text

Figure 1 from EST-SSR markers derived from an elite barley cultivar (Hordeum vulgare L. 'Morex'): polymorphism and genetic marker potential. | Semantic Scholar

Cultivar identification using seven selected SSR markers. a SSR allele... | Download Scientific Diagram

Development of EST–SSR markers in castor bean (Ricinus communis L.) and their utilization for genetic purity testing of hybrids
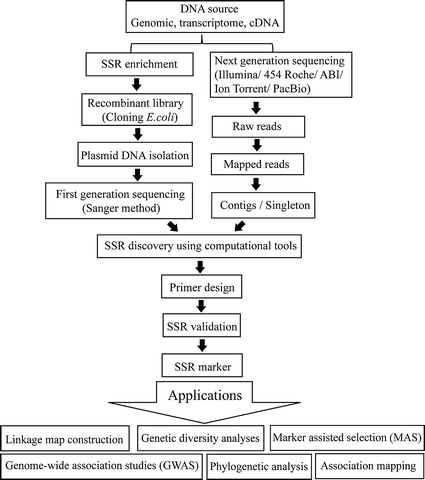
Simple sequence repeats (SSRs) markers in fish genomic research and their acceleration via next-generation sequencing and computational approaches | SpringerLink

Phylogenetic Analysis of 24 Brassica Species using SSR Markers By: Chavon T. Graves Alabama A & M REU Program 7/25/08 Fisk University. - ppt download

Diversity | Free Full-Text | SSR Markers for Trichoderma virens: Their Evaluation and Application to Identify and Quantify Root-Endophytic Strains | HTML